Create a gips object.
This object will contain initial data and all other information
needed to find the most likely invariant permutation.
It will not perform optimization. One must call
the find_MAP() function to do it. See the examples below.
Usage
gips(
S,
number_of_observations,
delta = 3,
D_matrix = NULL,
was_mean_estimated = TRUE,
perm = ""
)
new_gips(
list_of_gips_perm,
S,
number_of_observations,
delta,
D_matrix,
was_mean_estimated,
optimization_info
)
validate_gips(g)Arguments
- S
A matrix; empirical covariance matrix. When
Zis the observed data:if one does not know the theoretical mean and has to estimate it with the observed mean, use
S = cov(Z), and leave parameterwas_mean_estimated = TRUEas default;if one know the theoretical mean is 0, use
S = (t(Z) %*% Z) / number_of_observations, and set parameterwas_mean_estimated = FALSE.
- number_of_observations
A number of data points that
Sis based on.- delta
A number, hyper-parameter of a Bayesian model. It has to be strictly bigger than 1. See the Hyperparameters section below.
- D_matrix
Symmetric, positive-definite matrix of the same size as
S. Hyper-parameter of a Bayesian model. WhenNULL, the (hopefully) reasonable one is derived from the data. For more details, see the Hyperparameters section below.- was_mean_estimated
A boolean.
Set
TRUE(default) when yourSparameter is a result of astats::cov()function.Set FALSE when your
Sparameter is a result of a(t(Z) %*% Z) / number_of_observationscalculation.
- perm
An optional permutation to be the base for the
gipsobject. It can be of agips_permor apermutationclass, or anything the functionpermutations::permutation()can handle. It can also be of agipsclass, but it will be interpreted as the underlyinggips_perm.- list_of_gips_perm
A list with a single element of a
gips_permclass. The base object for thegipsobject.- optimization_info
For internal use only.
NULLor the list with information about the optimization process.- g
Object to be checked whether it is a proper object of a
gipsclass.
Value
gips() returns an object of
a gips class after the safety checks.
new_gips() returns an object of
a gips class without the safety checks.
validate_gips() returns its argument unchanged.
If the argument is not a proper element of a gips class,
it produces an error.
Functions
new_gips(): Constructor. It is only intended for low-level use.validate_gips(): Validator. It is only intended for low-level use.
Hyperparameters
We encourage the user to try D_matrix = d * I, where I is an identity matrix of a size
p x p and d > 0 for some different d.
When d is small compared to the data (e.g., d=0.1 * mean(diag(S))),
bigger structures will be found.
When d is big compared to the data (e.g., d=100 * mean(diag(S))),
the posterior distribution does not depend on the data.
Taking D_matrix = d * I is equivalent to setting S <- S / d.
The default for D_matrix is D_matrix = d * I, where
d = mean(diag(S)), which is equivalent to modifying S
so that the mean value on the diagonal is 1.
In the Bayesian model, the prior distribution for the covariance matrix is a generalized case of Wishart distribution.
For a brief introduction, see the Bayesian model selection
section in vignette("Theory", package = "gips") or in its
pkgdown page).
For analysis of the Hyperparameters influence, see Section 3.2.
of "Learning permutation symmetries with gips in R"
by gips developers Adam Chojecki, Paweł Morgen, and Bartosz Kołodziejek,
Journal of Statistical Software.
See also
stats::cov()- TheSparameter, as an empirical covariance matrix, is most of the time a result of thecov()function. For more information, see Wikipedia - Estimation of covariance matrices.find_MAP()- The function that finds the Maximum A Posteriori (MAP) Estimator for a givengipsobject.gips_perm()- The constructor of agips_permclass. Thegips_permobject is used as the base object for thegipsobject. To be more precise, the base object forgipsis a one-element list of agips_permobject.
Examples
require("MASS") # for mvrnorm()
perm_size <- 5
mu <- runif(5, -10, 10) # Assume we don't know the mean
sigma_matrix <- matrix(
data = c(
1.0, 0.8, 0.6, 0.6, 0.8,
0.8, 1.0, 0.8, 0.6, 0.6,
0.6, 0.8, 1.0, 0.8, 0.6,
0.6, 0.6, 0.8, 1.0, 0.8,
0.8, 0.6, 0.6, 0.8, 1.0
),
nrow = perm_size, byrow = TRUE
) # sigma_matrix is a matrix invariant under permutation (1,2,3,4,5)
number_of_observations <- 13
Z <- MASS::mvrnorm(number_of_observations, mu = mu, Sigma = sigma_matrix)
S <- cov(Z) # Assume we have to estimate the mean
g <- gips(S, number_of_observations)
g_map <- find_MAP(g, show_progress_bar = FALSE, optimizer = "brute_force")
g_map
#> The permutation (1,2,3,4,5):
#> - was found after 67 posteriori calculations;
#> - is 953.095 times more likely than the () permutation.
summary(g_map)
#> The optimized `gips` object.
#>
#> Permutation:
#> (1,2,3,4,5)
#>
#> Log_posteriori:
#> -0.5708509
#>
#> Times more likely than starting permutation:
#> 953.095
#>
#> The p-value of Likelihood-Ratio test:
#> 0.6008
#>
#> The number of observations:
#> 13
#>
#> The mean in the `S` matrix was estimated.
#> Therefore, one degree of freedom was lost.
#> There are 12 degrees of freedom left.
#>
#> n0:
#> 2
#>
#> The number of observations is bigger than n0 for this permutation,
#> so the gips model based on the found permutation does exist.
#>
#> The number of free parameters in the covariance matrix:
#> 3
#>
#> BIC:
#> 104.0469
#>
#> AIC:
#> 102.352
#>
#> --------------------------------------------------------------------------------
#> Optimization algorithm:
#> brute_force
#>
#> The number of log_posteriori calls:
#> 67
#>
#> Optimization time:
#> 0.1075909 secs
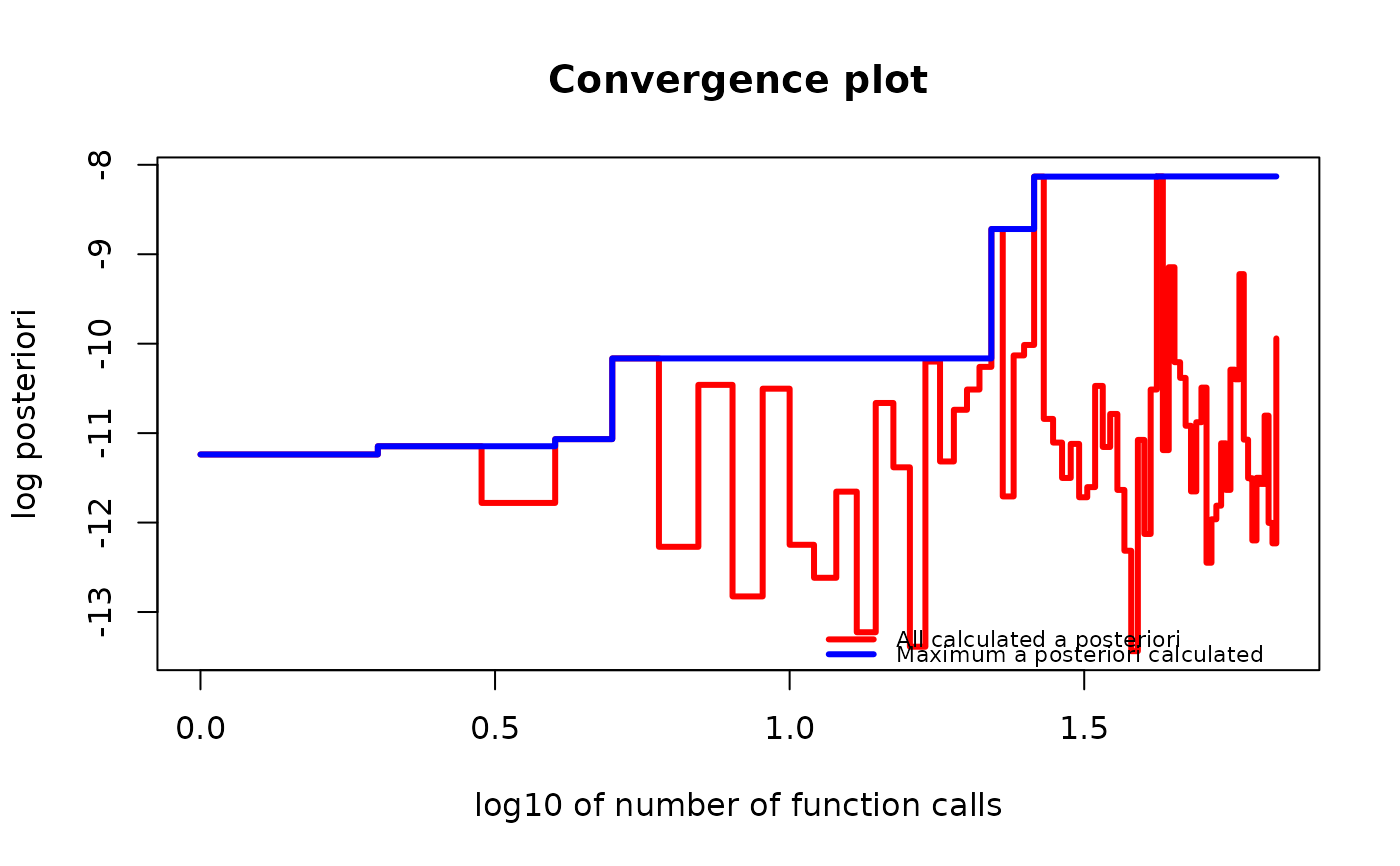
if (require("graphics")) {
plot(g_map, type = "both", logarithmic_x = TRUE)
}